Gene Prioritization with Word Embeddings and Protein Networks
Prioritizing candidate genes in ALS by integrating genetics with word embeddings.
🧬 Gene Prioritization with Word Embeddings and Protein Networks
📅 Research Period: Jul 2025 – Jul 2026
🌐 GitHub: Gene discovery
This research is supported by a FAPESP undergraduate fellowship (grant link).
Overview
Amyotrophic Lateral Sclerosis (ALS) is a fatal and highly heterogeneous neurodegenerative disease with multiple genetic and molecular contributors. Although GWAS and sequencing studies have identified important ALS-associated loci and genes, translating these signals into clear causal gene hypotheses remains a major challenge.
This project investigates whether biomedical literature embeddings can provide functional context for ALS gene prioritization. Instead of treating text-derived representations as standalone predictors, the project combines them with tissue-expression features from the Human Protein Atlas (HPA) and compares the resulting functional signal with GWAS-derived evidence.
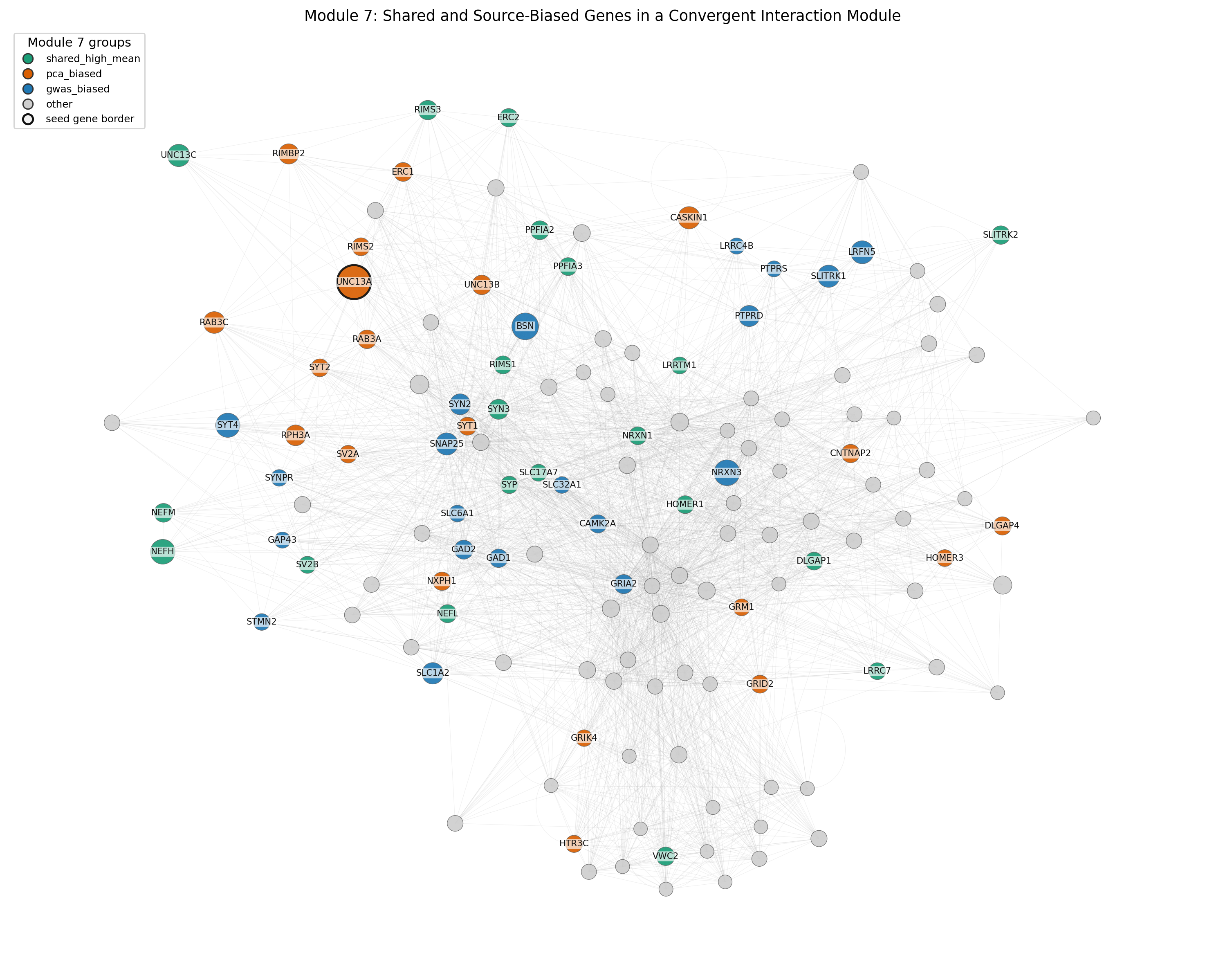
The main finding is that literature-derived embeddings did not consistently improve locus-level gene ranking, but the Word2Vec embedding space showed biologically interpretable structure. In protein network analyses, the Word2Vec/HPA model converged with GWAS-derived evidence at the level of protein interaction modules, suggesting that text-derived representations may be most useful for systems-level biological interpretation.
Method at a Glance
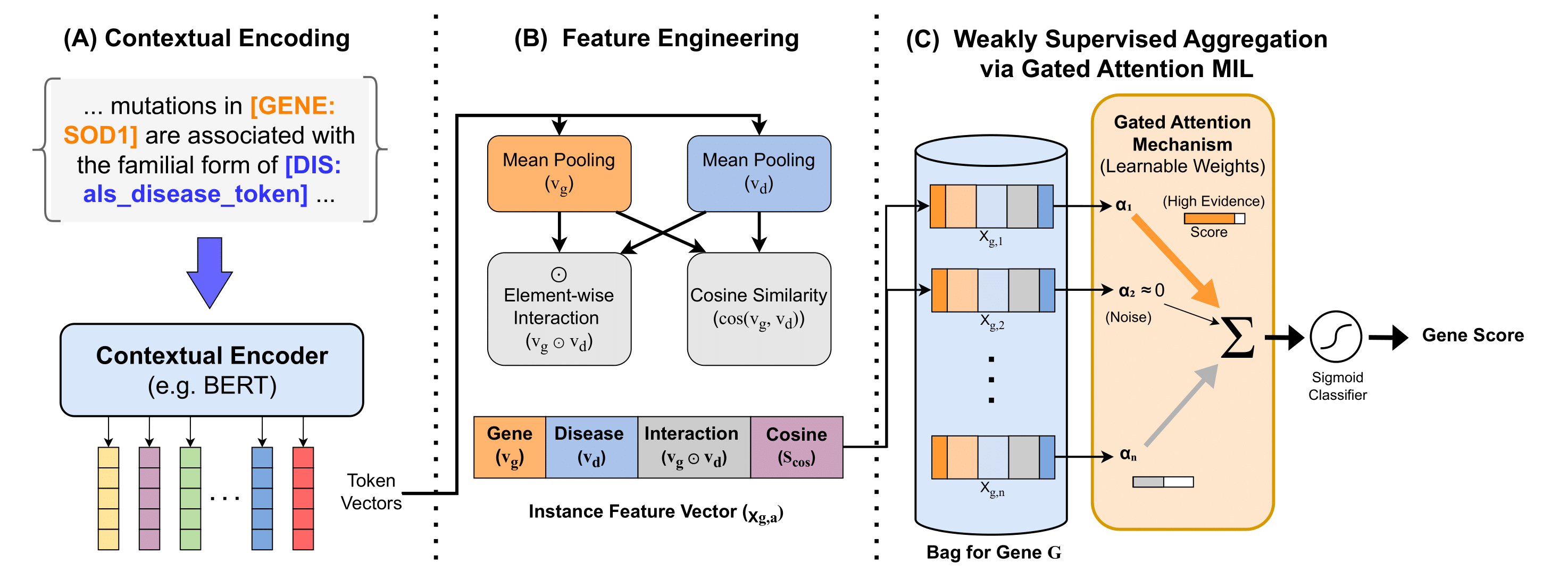
At a high level, the workflow consists of five stages:
- Literature collection: We construct disease-focused PubMed corpora for neurodegenerative and motor neuron disease contexts using ontology-guided search terms.
- Gene representation: Each gene is represented using literature-derived embeddings, with Word2Vec used as the main interpretable embedding model and PubMedBERT explored as a complementary contextual representation.
- Feature integration: Gene embeddings are reduced with PCA and combined with HPA brain and muscle expression features to build a functional gene-prioritization model.
- Locus-level evaluation: Logistic-regression models are evaluated on ALS-associated GWAS loci to test whether text-derived features improve candidate-gene ranking.
- Network interpretation: Functional scores from the Word2Vec/HPA model are compared with GWAS-derived evidence through protein-network propagation, module enrichment, and Gene Ontology analysis.